Testosterone and Frailty: A Decisive Shift Toward System-Level Biomarker Networks
[Ontological Limitations and the System-Level Mandate]
Conventional medical ontology predominantly conceptualizes hormonal biomarkers, such as testosterone, as solitary entities. Treatment decisions are often made based on local biological thresholds, treating similar biological levels as functionally identical without considering the surrounding system structure. However, this fragmented approach is insufficient for characterizing complex, high-order systemic states like frailty and aging. Physiological context matters fundamentally; similar testosterone levels can reflect fundamentally different underlying biological states depending on their interaction with dynamic and non-linear variables, including systemic inflammation, metabolic shifts, and declining organ function. This research addresses this limitation by proposing a definitive transformation of the medical ontology: the shift from viewing testosterone as an isolated variable, to conceptualizing it as an organized, high-dimensional, and structured relational system. Utilizing an immense real-world clinical dataset accumulated from high-volume, longitudinally followed surgical and medical practices, we apply high-order network science, discrete mathematics (including graph theory and topology), and data science techniques like unsupervised clustering. This integration enables us to model the intricate, discrete relational structures linking hormonal variables, inflammatory markers, renal and metabolic parameters, and diverse functional outcomes, facilitating a nuanced, system-level interpretation of physiological context.
[Unsupervised Clustering and Network Centrality]
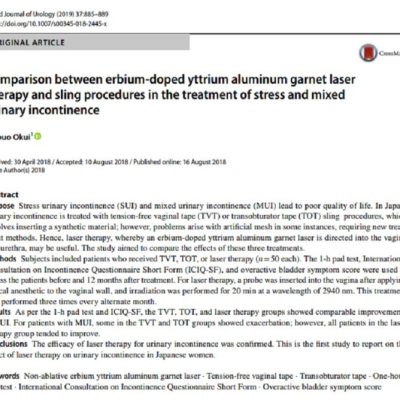
The methodological foundation of this research is a rigorous, quantitative transformation of multivariate data into structured, actionable networks. A key innovation of this work is the dynamic identification of higher-order topological patterns within these discrete relational models. Unsupervised clustering reveals distinct latent patient profiles characterized by non-linear combinations of biomarker variables. Our analysis has identified a striking high-risk aging profile, defined as a discrete relational motif where low testosterone coexists with systemic inflammation and impaired renal function. Importantly, this high-risk pattern is a coordinated biological state, not a simple hormonal abnormality. Network analysis further demonstrates that the mathematical importance (centrality) and relationship of testosterone are not fixed across individuals, but dynamically vary depending on the surrounding physiological context. Crucially, this research extends beyond male populations. Clinical studies in women show that testosterone is strongly associated with functional aging processes and urinary incontinence, particularly when integrated into a structured, high-dimensional network model. By treating these high-order network structures as dynamic mathematical objects, we can predict systemic response patterns and functional decline with unparalleled precision, reducing the trial-and-error approach common in traditional biomarker interpretation.
[Integrated Vision and Paradigmatic Transformation]
This visual and conceptual framework, illustrated as a bridge between a detailed narrative world and a high-order mathematical decision network, conveys a definitive paradigmatic shift. We are transforming reductionist clinical interpretation—which views hormones as disjointed biological signals—into a sophisticated, system-level understanding of biomarker structures. On the left side of this conceptual vision, traditional biomedical elements analyze organs independently. On the right, digital interfaces and artificial intelligence (AI) precisely represent the integration of real-world patient narratives and clinical data. The central triangular structure symbolizes the core innovation: the triadic decision motif that enables precise systemic characterization. By unifying clinical vocabulary, patient experiences, objective biological markers (hormonal, inflammatory, organ-specific), and dynamic anatomical markers into sophisticated relational models, Symptom Structure Science establishes a powerful new interface between quantitative precision and qualitative human suffering. This unifying framework enables a precise, data-driven interpretation of the true patient condition, facilitating the operationalization of personalized and truly patient-centered medicine. Ultimately, this work establishing a new framework of data-driven, system-level biomarker network Science, where treatment is guided not only by a single diagnosis, but by the underlying topological structure of a patient’s aging condition.